
Desk Report
Publish: 25 Jul 2021, 10:04 pm
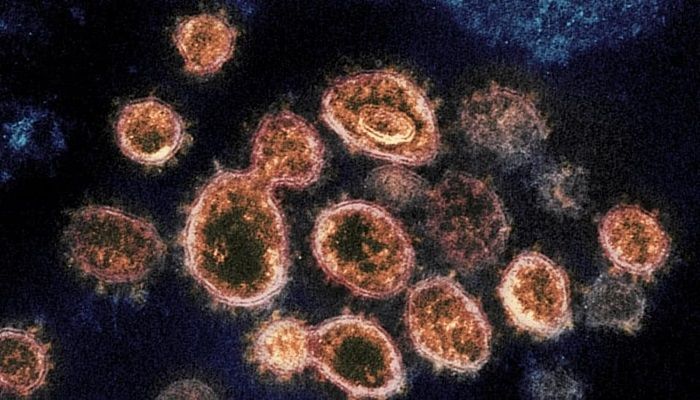
SARS-CoV-2 or Coronavirus (Photo: Collected)
A team of scientists, which researchers at the
University of Arizona in Tucson and the University of Adelaide in South
Australia co-led, delved into human genomes to find a correlation between
ancient coronavirus epidemics and past adaptation in modern humans.
They hope that understanding the effect of past
pandemics on genetic mutations will give scientists more “ammunition” in the
arms race against SARS-CoV-2 variants.
Yassine Souilmi, Ph.D., the lead author of the paper, is
a postdoctoral research associate at the Australian Centre for Ancient DNA. He
and his fellow researchers published their findings in the June 2021 edition of
Current Biology.
The authors explain in their article:
“Here, we apply evolutionary analyses to human genomic
datasets to recover selection events involving tens of human genes that
interact with coronaviruses, including SARS-CoV-2, that likely started more
than 20,000 years ago.”
VIPs and RNA viruses
Throughout human history, positive natural selection has
often targeted virus-interacting proteins (VIPs). VIPs either work in building
immunity or get hijacked by viruses.
This natural selection has persisted over the past
50,000 years, especially around VIPs that react to RNA viruses, such as
coronaviruses.
According to Souilmi and his team:
“The accumulated evidence suggests that ancient RNA
virus epidemics have occurred frequently during human evolution; however, we
currently do not know whether selection has made a substantial contribution to
the evolution of human genes that interact more specifically with
coronaviruses.”
Investigating VIP adaptations
Souilmi’s team culled genetic data from the 1000 Genomes
Project, a vast catalog of human genetic variations.
Two statistical analyses detected genetic signals known
as selective sweeps.
The researchers looked for selective sweep modifications
among more than 400 VIPs that interact with coronaviruses (CoV-VIPS). They
investigated data from across 26 populations.
Relying on evidence that VIPs are what viruses harness
to take over host cells, they focused on these genes. They also took this
approach because VIPs tend to exert a greater functional influence on viruses
compared with other proteins.
Sweep signals more than 900 generations
The scientists discovered a pattern of antiviral
modifications in sweep signals at 42 CoV-VIPs in five East Asian populations.
This enrichment did not appear in other populations.
The team reported:
“…[O]ur results are consistent with the emergence of a
viral epidemic… ∼25,000
years (28 years per generation) ago that drove a burst of strong positive
selection in East Asia. [These] selection events […] clearly predate the
estimated split of different East Asian populations included in the 1000
Genomes Project from their shared ancestral population.”
The observed mutations may have steadily increased in
frequency until about 200 generations, or an estimated 5,000 years, ago.
Also, the CoV-VIP proteins demonstrate antiviral and
proviral effects and variations that affect SARS-CoV-2 susceptibility and
COVID-19 severity in the current British population.
However, the paper notes that such adaptations in certain
human populations do not imply that those populations are more susceptible to
viral epidemics.
Medical News Today asked Martin Bachmann, Ph.D., an immunologist
and professor of vaccinology at the University of Oxford’s Jenner Institute in
the United Kingdom and the University of Bern in Switzerland, for his
perspective on this research:
“For me, it is quite stunning that you can analyze epidemics from 20,000 years ago without actually looking at a sample that is older than a few years. It shows that there is an awful lot of information buried in the genome of the whole population rather than individual genomes.”
Subscribe Shampratik Deshkal Youtube Channel
© 2024 Shampratik Deshkal All Rights Reserved. Design & Developed By Root Soft Bangladesh